-
Latest Version
SnapGene 8.2.2 LATEST
-
Review by
-
Operating System
Windows 7 / Windows 7 64 / Windows 8 / Windows 8 64 / Windows 10 / Windows 10 64
-
User Rating
Click to vote -
Author / Product
-
Filename
snapgene_8.2.2_win.exe
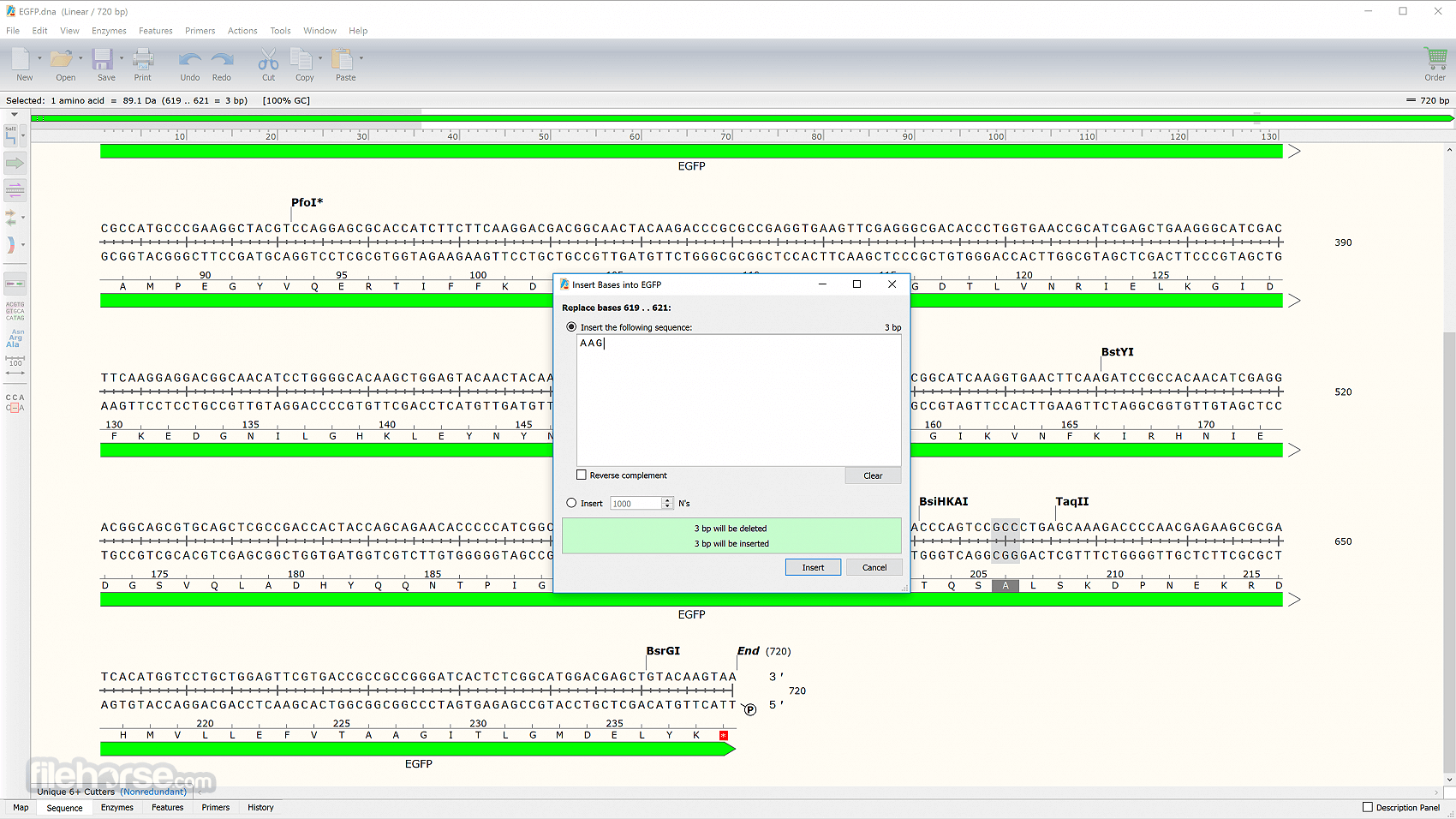
Take advantage of SnapGene’s efficient data handling to scan large DNA sequences with thousands of annotated features. Make insertions, deletions, replacements, and case changes. When a sequence is copied and pasted, features are automatically transferred. Annotate common features automatically, or annotate novel features manually.
Find common features in your DNA sequence using SnapGene’s extensive database. Additional features of your choosing can be added to a custom database.
It provides elegant, information-rich windows for simulating a variety of common cloning and PCR methods. Highlight unique restriction sites in bold font, or choose the automatically defined Unique Cutters or Unique 6+ Cutters enzyme set.
Use your own primers, or ask the app to design primers automatically. The product file stores the template and primers in its history. Assemble up to eight fragments. Select the fragments to be joined and their orientations, and Snap Gene will design primers.
Use a Sequence view to see at a glance whether two translated features are in the frame. If so, the translations are linked on the same line. If not, the translations are on separate lines. Use the powerful alignment tool to check whether an actual construct matches the simulated construct.
It automatically records operations to create a graphical history, and stores the ancestor constructs in the final file. Use the familiar, secure operating system of your computer to store and organize your Snap Gene files.
Export a sequence to GenBank or FASTA format. Export a map or simulated agarose gel to common image formats. Convert a sequence, map, or gel image to standard formats for use with other software.

The open exchange of information is crucial, so SnapGene and SnapGene Viewer provide options for reading and exporting common file formats.
How to Use
- Download SnapGene from the official website
- Install the software on your Windows PC
- Launch the application after installation
- Create or open a DNA sequence file
- Annotate features and edit sequences
- Simulate cloning procedures
- View plasmid maps and sequence details
- Export data or save your project
- Share files with collaborators
- Use tutorials for advanced features

Pricing
Academic Subscription: For universities and nonprofits using SnapGene for academic research. Annual pricing depends on the number of users — $350 for 1 user, $650 for 2, $1,625 for 5, $3,000 for 10. Includes all platform licenses, updates, and license management tools.
Corporate Subscription: For commercial use, priced higher — $1,845 for 1 user, $3,690 for 2, $9,225 for 5, $18,450 for 10 users per year. Offers the same benefits as the academic plan but for business environments.
Student Subscription: Available to enrolled students (undergrad, grad, postdoc, med residents) for $149/year. Must be purchased online by the individual student. Intended for academic work only.
Permanent Licenses: One-time purchase option with no updates or support. $2,500 for academic users and $9,600 for corporate. Not recommended due to lack of upgrades or compatibility guarantees.
Course Licenses: Free temporary licenses for educators offering SnapGene to students as part of a class.
All paid plans support installation on Windows, Mac, and Linux, and offer user management, deployment tools, and activation control. Corporate and Academic plans also support prorated additions and automatic renewal.

System Requirements
Operating System: Windows 11 or Windows 10 (64-bit)
Processor: 2.4 GHz or faster
RAM: Minimum 4 GB (8 GB recommended)
Hard Disk Space: At least 500 MB available
Display: 1280x800 resolution or higher
Internet: Required for activation and updates
PROS
- User-friendly interface
- Accurate cloning simulation
- Real-time plasmid visualization
- Broad file format support
- Extensive annotation tools
- Expensive licensing cost
- Limited free version features
- No native Linux support
- High resource consumption
- Steep learning curve for some users
What's new in this version:
Changed:
- Improved Japanese and Chinese translations
- Simplified the command line usage message
- Updated information for restriction enzymes BSmFI, BshVI, and BSSHII
- Replaced the Y-FAST feature with an updated FAST feature in the common features database
- Updated list of enzymes provided by Jena Bioscience
Fixed:
- Improved stability of circular sequence maps
- Fixed a crash in Golden Gate cloning that occurred when flipping the insert twice
- Fixed a crash when interacting with files in the project side panel
- Fixed a crash on startup when launching SnapGene from the macOS Dock
- Fixed pointer ghosting when interacting with reads aligned to a reference sequence
- Fixed ORF boundary detection when the stop codon spans the origin
 OperaOpera 131.0 Build 5877.24 (64-bit)
OperaOpera 131.0 Build 5877.24 (64-bit) Kling AIKling AI - Text or Image to Video
Kling AIKling AI - Text or Image to Video PhotoshopAdobe Photoshop CC 2026 27.5 (64-bit)
PhotoshopAdobe Photoshop CC 2026 27.5 (64-bit) BlueStacks AIBlueStacks AI
BlueStacks AIBlueStacks AI OKXOKX - Buy Bitcoin or Ethereum
OKXOKX - Buy Bitcoin or Ethereum CapCutCapCut 8.5.0
CapCutCapCut 8.5.0 PC RepairPC Repair Tool 2026
PC RepairPC Repair Tool 2026 Hero WarsHero Wars - Online Action Game
Hero WarsHero Wars - Online Action Game TradingViewTradingView - Trusted by 100 Million Traders
TradingViewTradingView - Trusted by 100 Million Traders AdGuard VPNAdGuard VPN 2.9.0
AdGuard VPNAdGuard VPN 2.9.0






Comments and User Reviews